Dataframe interchange protocol and Vaex
The work I briefly describe in this blog post is the implementation of the dataframe interchange protocol into Vaex which I was working on through the three month period as a Quansight Labs Intern.
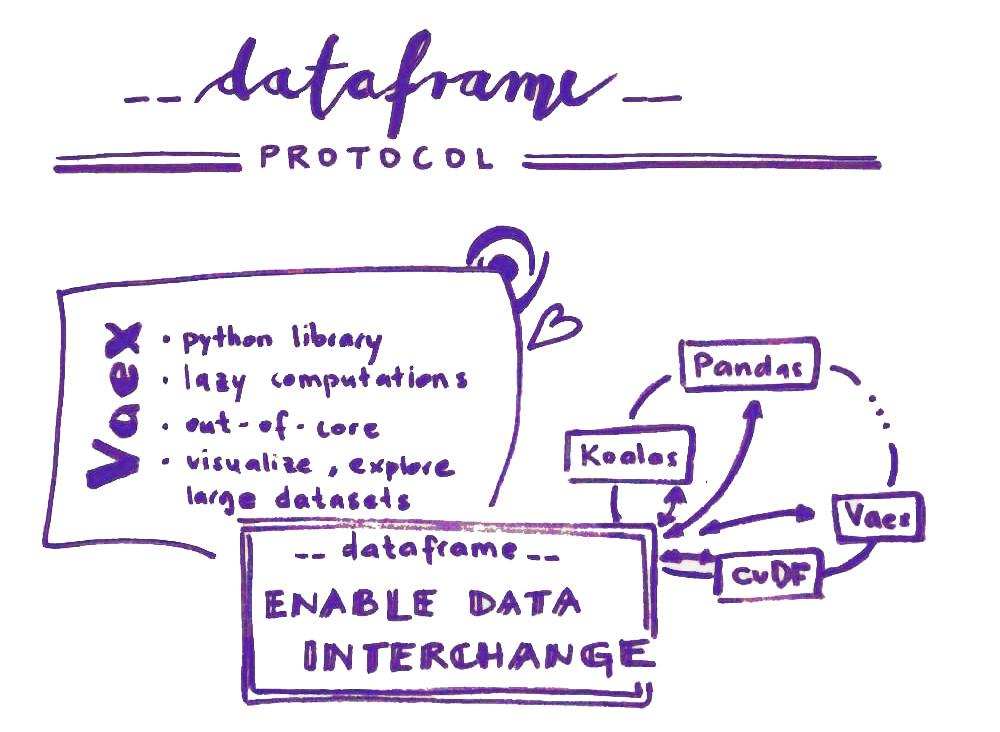 Connection between dataframe libraries with dataframe protocol
Connection between dataframe libraries with dataframe protocol
About | What is all that?
Today there are quite a number of different dataframe libraries available in Python. Also, there are quite a number of, for example, plotting libraries. In most cases they accept only the general Pandas dataframe and so the user is quite often made to convert between dataframes in order to be able to use the functionalities of a specific plotting library. It would be extremely cool to be able to use plotting libraries on any kind of dataframe, would it not?
Dataframe protocol
The purpose of the Dataframe interchange protocol (
__dataframe__) is to enable data interchange. I.e., a way to convert one type of dataframe into another type (for example, convert a Koalas dataframe into a Pandas dataframe, or a cuDF dataframe into a Vaex dataframe).
With the protocol implemented in dataframe libraries we will be able to write code that accepts any kind of dataframe 🎉
For more information about the protocol visit the RFC blog post or the official site.
Dataframe
Besides the possibility of converting one type of dataframe into another the design of the protocol enables us to inspect the dataframe for basic properties like the number of columns, missing values, etc. At this point we could ask ourselves what is the general definition of the dataframe?
For the purpose of the dataframe protocol the term is defined as follows:
A dataframe is an ordered collection of columns, which are conceptually 1-D arrays with a dtype and missing data support. A column has a name, which is a unique string. A dataframe or a column may be “chunked”, meaning its data is not contiguous in memory.
Vaex
One of the dataframe libraries that is used very often for its high performance is Vaex.
Vaex is a high performance Python library for lazy Out-of-Core DataFrames, to visualize and explore big tabular datasets. It can calculate statistics such as mean, sum, count, standard deviation etc, on an N-dimensional grid up to a billion objects/rows per second. Visualization is done using histograms, density plots and 3d volume rendering, allowing interactive exploration of big data. Vaex uses memory mapping, a zero memory copy policy, and lazy computations for best performance (no memory wasted).
More about Vaex is available on the official site and blog.
Vaex has many great features! With the implementation of the dataframe protocol other libraries such as Plotly could accept a Vaex dataframe and so the functionalities of both libraries would be enhanced.
Implementation
Implementation of the dataframe protocol connects the Vaex dataframe class with the __dataframe__ method specified by the Consortium for Python Data API Standards. It also defines the constructor function called from_dataframe with which the conversion between dataframes is made.
The __dataframe__ method returns an instance of the _DataFrame class. At this point I would like to quickly describe the conceptual model of how the data is represented in memory. After that I will go through the steps I took to implement the protocol for Vaex.
The smallest building blocks in the memory representation of the dataframe are 1-D arrays (or "buffers"), which are contiguous in memory and contain data with the same dtype. A column consists of one or more 1-D arrays (if, e.g., missing data is represented with a boolean mask, that's a separate array). A dataframe contains one or more columns. A column or a dataframe can be "chunked"; a chunk is a subset of a column or dataframe that contains a set of (neighboring) rows.
.](../../../../images/2021/09/dataframe-api-vaex/dataframe-api-vaex_UML.jpg) UML diagram
UML diagram
For the memory representation of the dataframe three separate classes are defined. These are _Buffer, _Column and _DataFrame. In the Vaex implementation we named them _VaexBuffer, _VaexColumn and _VaexDataFrame respectively. Each of them has necessary and utility methods to construct and describe a dataframe.
The constructor function from_dataframe iterates through the chunks and through the dictionary of columns of the input dataframe, calls the correct methods and converts the column to the desired type.
With the general overview of the protocol and its model in mind I will next present the path I took to implement it into Vaex.
Implementation steps | Interesting topics covered, interesting lessons learned
How did I start?
First thing I did was reading and re-reading the specifications. After that I talked about the problem as much as possible. With my mentor, with my colleague, with a friend, ... Talking about it helped me get a bigger picture and an idea of where to start. Starting is hard. Talking about it helps ☕️💗
I had to make clear which dtypes the protocol needs to support and how I can handle them in Vaex. With that I could start with simple dtypes and work my way to the ones hardest to implement.
To find out more about data types in the dataframe protocol visit this DataFrame API issue on GitHub.
A prototype implementation for Pandas dataframe had already been written so I started here with the most simple part - int and float dtypes. Very soon in the process of researching I had to get acquainted with the term buffers and array interface.
Array interface and buffers
When the from_dataframe method iterates through the columns it basically transfers the problem to the columnar level where the interchange of data is really happening. That means array API is used.
Array API? Similar to the idea of converting one type of dataframe into another, array API interchange mechanism supports converting one type of array into another. An array is a (usually fixed-size) multidimensional container of items of the same type and size (source). You can also call it a column, tensor, ... For more information see the Data interchange mechanisms section in the Data APIs documentation.
There are three relevant existing protocols that support array interchange and they are buffer protocol, __array_interface__ and DLPack.
The best candidate for the array API is chosen to be DLPack. Pandas and Vaex don't have this protocol implemented so instead we receive the information about the location where the data is stored in memory with the use of array_interface. Therefore in Pandas and Vaex implementations the columns are read as NumPy arrays (ndarrays) via __array_interface__.
The specifications of the buffer, for example, buffer size, a pointer to the start of the buffer, DLPack attributes and a string representation of the buffer, are a part of the _VaexBuffer class.
In the _VaexColumn class the data type description is added in .dtype as a tuple (kind, bit-width, format string, endianness). It is used as a descriptive attribute and also as an input to construct a NumPy array from the buffer. The implementation of the dtype got more complicated when dealing with categoricals and string dtypes.
The function from the _VaexColumn class that saves the data into a _VaexBuffer class to be converted in the process is get_buffers. It returns a dictionary of three separate two-element tuples:
-
_get_data_buffer: buffer containing the data and its associated dtype, -
_get_validity_buffer: buffer containing mask values indicating missing data and its associated dtype, -
_get_offsets_buffer: offset values for variable-size binary data (e.g., variable-length strings) and its associated dtype.
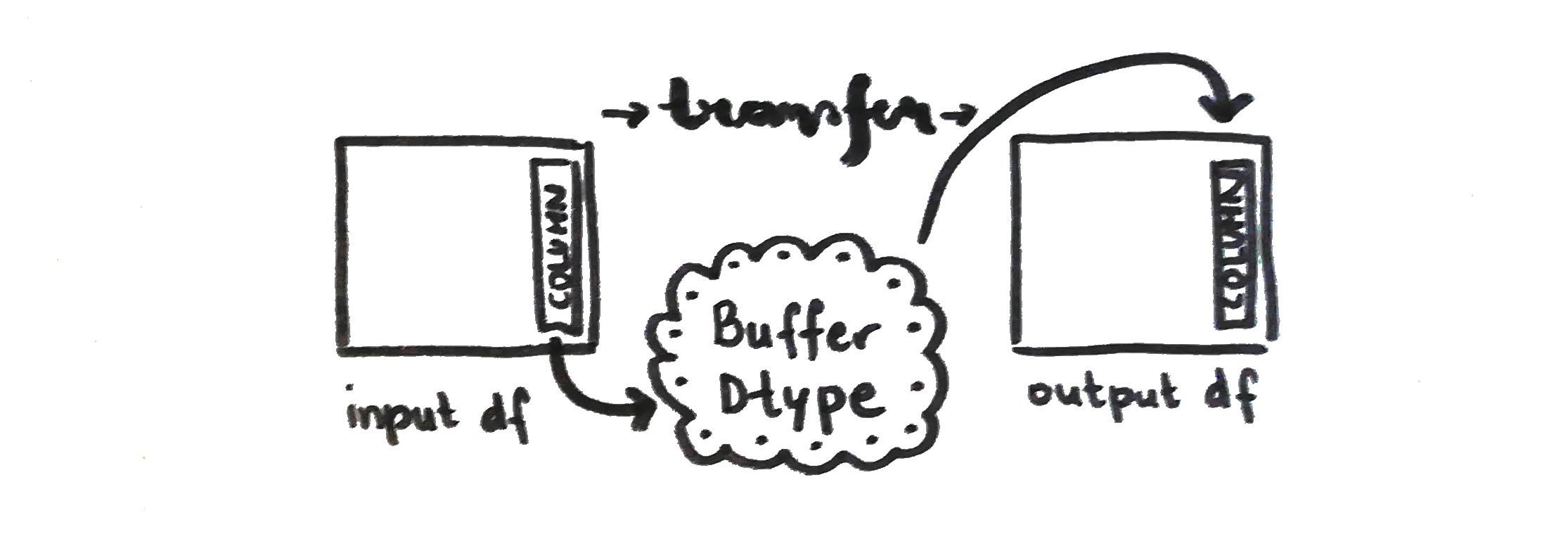
This is used in the from_dataframe method when columns are iterated through. The method calls the get_buffers function and converts the data via __array_interface__. If, for example, the user wants to convert Pandas dataframe to a Vaex instance, the Vaex from_dataframe method (that we called from_dataframe_to_vaex just to make it clearer) calls Pandas get_buffers method and then makes the conversion.
 Buffers
Buffers
Understanding the conversion cycle for simple dataframe dtypes by following Pandas implementation outline helped me implement the protocol for Vaex and after the tests were not giving any errors I added a boolean column which worked fine. The next stop was categorical dtypes.
Categoricals
In this case the codes of the categories are the ones being converted through the buffer. Categories are mapped and than applied to the codes afterwords. The function from _VaexColumn class used to determine the mapping is describe_categorical. It returns a dictionary with a boolean value indicating whether the ordering of dictionary indices is semantically meaningful, a second boolean value indicating whether a dictionary-style mapping of categorical values to other objects exists and finally the mapping of the categories.
Based on my research you can have Vaex categorical columns made with methods categorize or ordinal_encode (which is deprecated). There is another possibility where the underlying expression is an Arrow dictionary. Both options have different functions to be used.
Additionally there is a special case in Veax when categorize function is used and codes need to be calculated separately (year column in the example).
Categoricals are complicated. =) I was happy when the tests finally passed.
Missing values
Clarifying what a missing value should and should not be is quite difficult and there isn't one way of looking at it. True missing and NaN are different things altogether but when analysing you deal with unavailable data in both cases. The other question is how to store missing values? Nullable, sentinel, with bit or byte mask,...?
More about this topic can be found in this issue.
Methods in the _VaexColumn class used for the missing data are the number of missing values null_count and describe_null which returns the missing value (or "null") representation the column dtype uses, as a tuple (kind, value). "Kind" can be non-nullable, NaN/NaT, sentinel value, bit mask or byte mask. The "value" can be the actual (sentinel) value, (0 or 1) indicating a missing value in case of a mask representation or None otherwise.
All the missing values in Vaex are nullable. Arrow columns via bitmask, NumPy columns via bool/byte arrays. For defining a validity buffer I used a boolean array returned from Vaex's .ismissing() method and worked with that.
The method from_dataframe needs to check for the null representation and apply the missing values accordingly. In the implementation for Vaex the validity buffer is taken as a mask input for the Arrow array with which the transformed data is constructed into an array/column. In case of sentinel values the value from describe_null is used to construct a mask and use it in the Arrow array also.
Virtual columns
Another great thing about Vaex! As it lazily evaluates the data a newly defined column/expression will not be saved to memory unless specified. This kind of en expression/column is called a virtual column. As far as the implementation is concerned the protocol materializes them and handles them as normal columns.
So no additional work was needed here.
Next in line were chunks.
Chunks
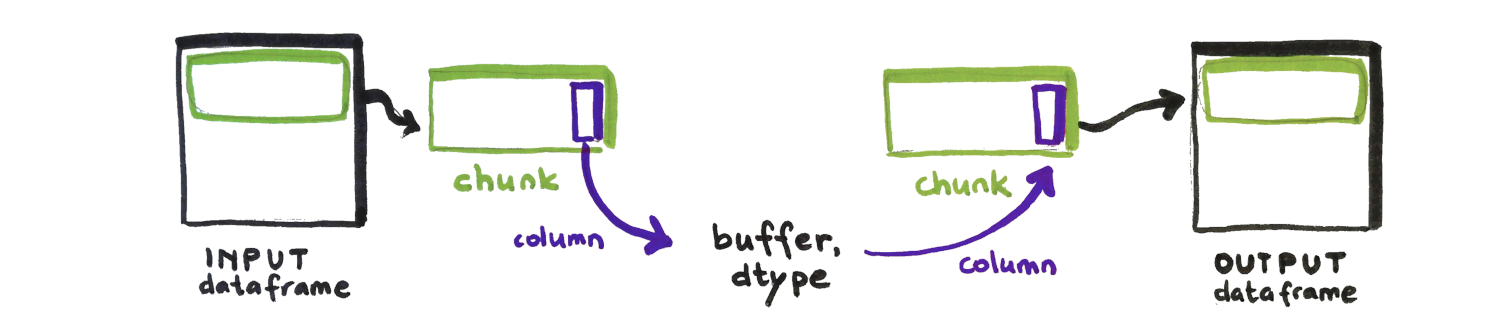
A chunk is a subset of a column or dataframe that contains a set of (neighboring) rows.
A nice visual explaining chunks can be seen here.
The _VaexColumn and _VaexDataFrame methods used in the case of chunked dataframe are num_chunks and get_chunks. With the former, the from_dataframe method checks if the dataframe is chunked or not and iterates through the chunks if necessary. The iterator is the output of the get_chunks method from the _VaexDataFrame class.
Vaex can read data in chunks. It is not the only library that does so and it is a functionality that should be preserved if possible.
 Chunks
Chunks
String dtype
The last thing that was implemented as part of my internship was the handling of variable-length string dtypes. The Pandas implementation already had a great PR merged and it turned out it could easily be applied to Vaex as well.
In the get_buffers method the data needs to be encoded to pass it through the buffer. Also the offset of each string is saved to the offset buffer in order to be able to decode the strings when converting from buffers in from_dataframe.
In all the above steps the effort of implementing the protocol wasn't just mine, it was a joint endeavor of the developers from Quansight and the Vaex team. The help received from both ends enabled me to progress and to find better solutions.
Journey through the internship
 My three-month journey
My three-month journey
I will end this blog post with a little bit of a personal note.
For me the work at Quansight in the least three months meant lots of space to figure things out. I don't have trouble getting things done. If, with this freedom to work, you can also receive the right amount of interaction, motivation, community vibe, ... that is just awesome. And that was how I experienced the internship.
My mentor, Kshiteej Kalambarkar, was always available for a technical question or just a chat. He helped me with on-boarding and with finalizing the work. My co-intern Ismaël was always ready to talk things over. He helped me with lots of silly things that were blocking me.
The onboarding process was very friendly with a perfect rhythm. Most of the information about the process passed through email. The Labs Internship Handbook was circulated containing all possible info you might need. We received a warm welcome via Slack and from day one on we had a weekly Intern Share where all the interns got together. At first to introduce ourselves to each other, later to present, hear about and support the work in progress. Presentations were interesting from a knowledge as well as a social perspective.
Connecting with the developers
Thanks to Ralf Gommers for getting us connected with the Vaex developers. They are extremely friendly, understanding and motivated. Always ready to help. After submitting a draft PR, Maarten Breddels and Jovan Veljanoski helped a lot and my work got pushed further.
What I found very helpful in the process was making a personal repository where I saved my ongoing work and made issues to track the working process. In this way I was able to organize better and the stress of not knowing what exactly should be done next or how fast I should be working was reduced. Also the sharing of work with colleagues and mentor was easier.
At the end the experience as a Quansight Intern was highly positive and I am planning to stay engaged in the work of the dataframe API protocol even when the internship is over.
Example Notebook
If the topic is of interest to you, here is an Example Notebook you can try out with a live Python kernel:
Thank you for reading through.
 Thank you!
Thank you!
Comments